These functions can be used to generate null distributions for testing the prevalence of imaginary CSS. The null is a function of the individual level accuracy rates, in other words. Pr(i perceives a one| there is a one) and Pr(i perceives a zero| there is a zero).
Usage
tie_level_accuracy(graph, which_nets = NULL)
sample_css_network(
graph,
prob = tie_level_accuracy(graph),
i = 1L:attr(graph, "netsize"),
keep_baseline = TRUE
)Arguments
- graph
A barry_graph object.
- which_nets
Integer vector. The networks to sample from.
- prob
A numeric vector of length 4L or a data frame (see details).
- i
Integer vector. The network to sample from.
- keep_baseline
Logical scalar. When
TRUE, the function returns the baseline network as the first element of the list.
Value
The function tie_level_accuracy returns a data frame with the following columns:
k: The perceiver id.p_0_ego: The probability of no tie between the perceiver (ego) and an alter.p_1_ego: The probability of a tie between the perceiver and an alter.p_0_alter: The probability of no tie between two alters.p_1_alter: The probability of a tie between two alters.
The function sample_css_network returns a list of square matrices of size
attr(graph, "netsize"). If keep_baseline = TRUE, the first element of
the list is the baseline network. Otherwise, it is not returned.
Details
There are two special cases worth mentioning. First, when the dyads in
question are all present the probability of true negative is set to NA.
On the other hand, if the dyads in question are all null, the probability of
true positive is NA as well. This doesn't affect the sample_css_network
function because those probabilities are unsed since tie/no tie probabilities
are according to the baseline graph, meaning that, for instance, a fully
connected network will never use the p_0_ego and p_0_alter
probabilities and an empty network will never use the p_1_ego and
p_1_alter probabilities.
The function sample_css_network samples perceived networks from the
baseline network. The baseline network is the first network in the
graph object. The function tie_level_accuracy can be used to
generate the probability vector.
The probability vector is a numeric vector of length 4L. The first
two elements are the probability of a tie/no tie between an ego and an alter.
The third and fourth elements are the probability of a tie/no tie between
two alters. When prob is a data frame, the function will sample from
each row of the data frame (returned from the function tie_level_accuracy).
Examples
# Create example networks
true_net <- matrix(c(0, 1, 1, 0,
1, 0, 0, 1,
1, 0, 0, 1,
0, 1, 1, 0), nrow = 4, byrow = TRUE)
# Person 1's perception (some errors)
person1_view <- matrix(c(0, 1, 0, 0,
1, 0, 1, 1,
0, 1, 0, 0,
0, 1, 0, 0), nrow = 4, byrow = TRUE)
# Person 2's perception (different errors)
person2_view <- matrix(c(0, 1, 1, 1,
1, 0, 0, 0,
1, 0, 0, 1,
1, 0, 1, 0), nrow = 4, byrow = TRUE)
# Create barry graph
networks <- list(true_net, person1_view, person2_view)
graph <- new_barry_graph(networks)
# Calculate accuracy rates
accuracy <- tie_level_accuracy(graph)
print(accuracy)
#> k p_0_ego p_1_ego p_0_alter p_1_alter
#> 1 1 1 0.5 0 0.0
#> 2 2 1 0.5 1 0.5
# Visualize accuracy patterns
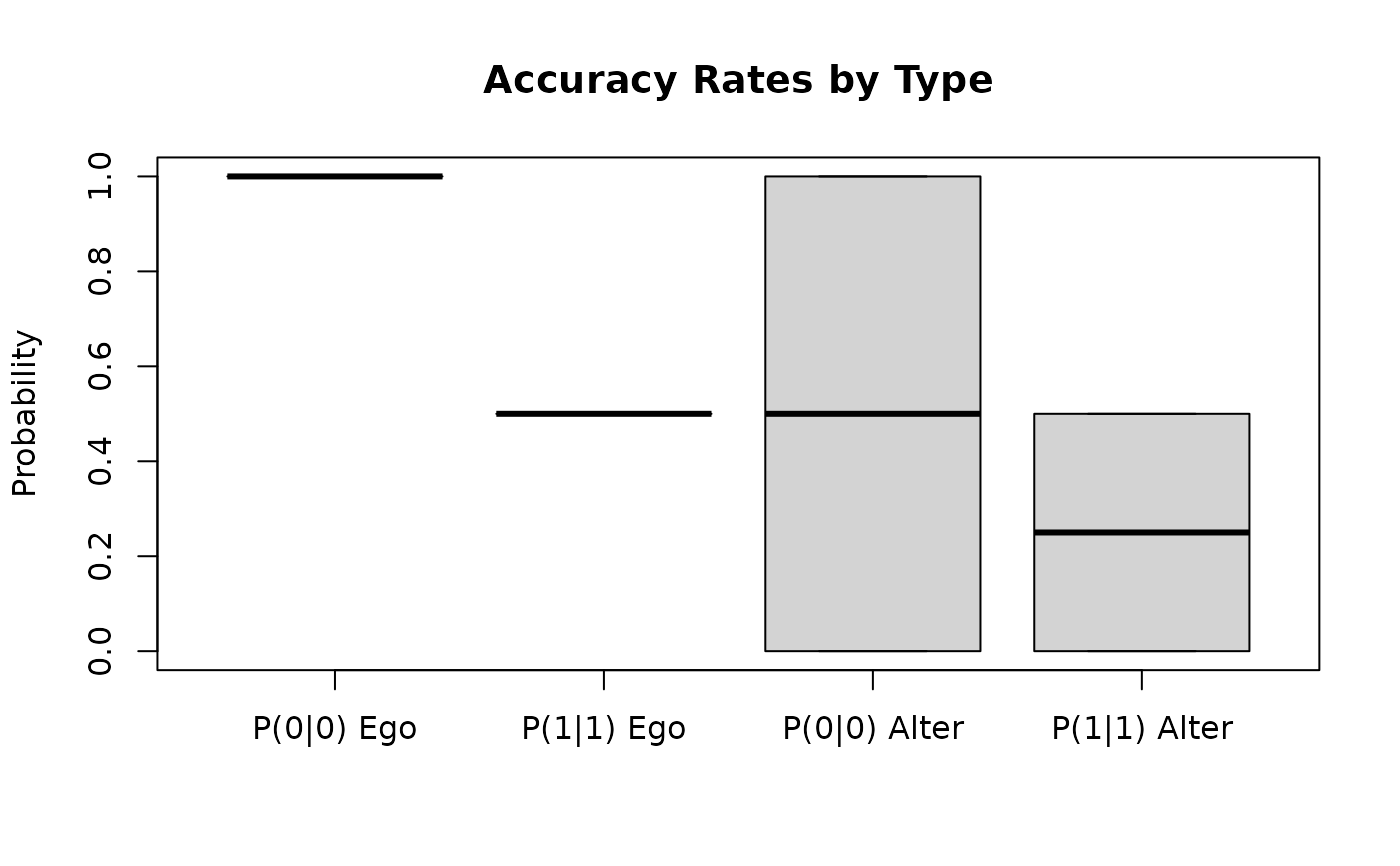
boxplot(accuracy[, -1],
main = "Accuracy Rates by Type",
ylab = "Probability",
names = c("P(0|0) Ego", "P(1|1) Ego", "P(0|0) Alter", "P(1|1) Alter"))
 # Calculate for specific networks only
accuracy_subset <- tie_level_accuracy(graph, which_nets = c(1, 2))
print(accuracy_subset)
#> k p_0_ego p_1_ego p_0_alter p_1_alter
#> 1 1 1 0.5 0 0.0
#> 2 2 1 0.5 1 0.5
# Create example networks for sampling
baseline <- matrix(c(0, 1, 1, 0,
1, 0, 0, 1,
1, 0, 0, 1,
0, 1, 1, 0), nrow = 4, byrow = TRUE)
perception1 <- matrix(c(0, 1, 0, 0,
1, 0, 1, 1,
0, 1, 0, 0,
0, 1, 0, 0), nrow = 4, byrow = TRUE)
perception2 <- matrix(c(0, 1, 1, 1,
1, 0, 0, 0,
1, 0, 0, 1,
1, 0, 1, 0), nrow = 4, byrow = TRUE)
# Create graph
graph <- new_barry_graph(list(baseline, perception1, perception2))
# Method 1: Using accuracy data frame (recommended)
accuracy <- tie_level_accuracy(graph)
sampled_networks <- sample_css_network(graph, prob = accuracy)
# Print number of networks generated
cat("Generated", length(sampled_networks), "networks\n")
#> Generated 3 networks
# Check first sampled network
print("First sampled network:")
#> [1] "First sampled network:"
print(sampled_networks[[2]]) # [[1]] is baseline if keep_baseline=TRUE
#> [,1] [,2] [,3] [,4]
#> [1,] 0 1 0 0
#> [2,] 1 0 1 0
#> [3,] 1 1 0 0
#> [4,] 0 0 0 0
#> attr(,"probs")
#> [,1] [,2] [,3] [,4]
#> [1,] NA 0.5 0.5 0
#> [2,] 0.5 NA 1.0 0
#> [3,] 0.5 1.0 NA 0
#> [4,] 0.0 0.0 0.0 NA
# Method 2: Using manual probability vector
# p_0_ego, p_1_ego, p_0_alter, p_1_alter
manual_probs <- c(0.8, 0.9, 0.85, 0.75)
sampled_manual <- sample_css_network(
graph,
prob = manual_probs,
i = 1,
keep_baseline = FALSE
)
print("Manual probability sampling:")
#> [1] "Manual probability sampling:"
print(sampled_manual[[1]])
#> NULL
# Method 3: Generate multiple samples for simulation
n_simulations <- 5
simulation_results <- replicate(n_simulations, {
sample_css_network(graph, prob = accuracy, keep_baseline = FALSE)
}, simplify = FALSE)
cat("Generated", length(simulation_results), "simulation rounds\n")
#> Generated 5 simulation rounds
cat("Each round has", length(simulation_results[[1]]), "networks\n")
#> Each round has 2 networks
# Compare original vs sampled network properties
original_density <- mean(baseline)
sampled_density <- mean(sampled_networks[[2]])
cat("Original network density:", round(original_density, 3), "\n")
#> Original network density: 0.5
cat("Sampled network density:", round(sampled_density, 3), "\n")
#> Sampled network density: 0.312
# Calculate for specific networks only
accuracy_subset <- tie_level_accuracy(graph, which_nets = c(1, 2))
print(accuracy_subset)
#> k p_0_ego p_1_ego p_0_alter p_1_alter
#> 1 1 1 0.5 0 0.0
#> 2 2 1 0.5 1 0.5
# Create example networks for sampling
baseline <- matrix(c(0, 1, 1, 0,
1, 0, 0, 1,
1, 0, 0, 1,
0, 1, 1, 0), nrow = 4, byrow = TRUE)
perception1 <- matrix(c(0, 1, 0, 0,
1, 0, 1, 1,
0, 1, 0, 0,
0, 1, 0, 0), nrow = 4, byrow = TRUE)
perception2 <- matrix(c(0, 1, 1, 1,
1, 0, 0, 0,
1, 0, 0, 1,
1, 0, 1, 0), nrow = 4, byrow = TRUE)
# Create graph
graph <- new_barry_graph(list(baseline, perception1, perception2))
# Method 1: Using accuracy data frame (recommended)
accuracy <- tie_level_accuracy(graph)
sampled_networks <- sample_css_network(graph, prob = accuracy)
# Print number of networks generated
cat("Generated", length(sampled_networks), "networks\n")
#> Generated 3 networks
# Check first sampled network
print("First sampled network:")
#> [1] "First sampled network:"
print(sampled_networks[[2]]) # [[1]] is baseline if keep_baseline=TRUE
#> [,1] [,2] [,3] [,4]
#> [1,] 0 1 0 0
#> [2,] 1 0 1 0
#> [3,] 1 1 0 0
#> [4,] 0 0 0 0
#> attr(,"probs")
#> [,1] [,2] [,3] [,4]
#> [1,] NA 0.5 0.5 0
#> [2,] 0.5 NA 1.0 0
#> [3,] 0.5 1.0 NA 0
#> [4,] 0.0 0.0 0.0 NA
# Method 2: Using manual probability vector
# p_0_ego, p_1_ego, p_0_alter, p_1_alter
manual_probs <- c(0.8, 0.9, 0.85, 0.75)
sampled_manual <- sample_css_network(
graph,
prob = manual_probs,
i = 1,
keep_baseline = FALSE
)
print("Manual probability sampling:")
#> [1] "Manual probability sampling:"
print(sampled_manual[[1]])
#> NULL
# Method 3: Generate multiple samples for simulation
n_simulations <- 5
simulation_results <- replicate(n_simulations, {
sample_css_network(graph, prob = accuracy, keep_baseline = FALSE)
}, simplify = FALSE)
cat("Generated", length(simulation_results), "simulation rounds\n")
#> Generated 5 simulation rounds
cat("Each round has", length(simulation_results[[1]]), "networks\n")
#> Each round has 2 networks
# Compare original vs sampled network properties
original_density <- mean(baseline)
sampled_density <- mean(sampled_networks[[2]])
cat("Original network density:", round(original_density, 3), "\n")
#> Original network density: 0.5
cat("Sampled network density:", round(sampled_density, 3), "\n")
#> Sampled network density: 0.312